Spark is a platform for cluster computing.
- Spark lets you spread data and computations over clusters with multiple nodes (think of each node as a separate computer).
- Splitting up your data makes it easier to work with very large datasets because each node only works with a small amount of data.
- As each node works on its own subset of the total data, it also carries out a part of the total calculations required, so that both data processing and computation are performed in parallel over the nodes in the cluster.
- It is a fact that parallel computation can make certain types of programming tasks much faster.
How to decide if you want to use Spark?
- Is my data too big to work with on a single machine?
- Can my calculations be easily parallelized?
Using Spark in Python
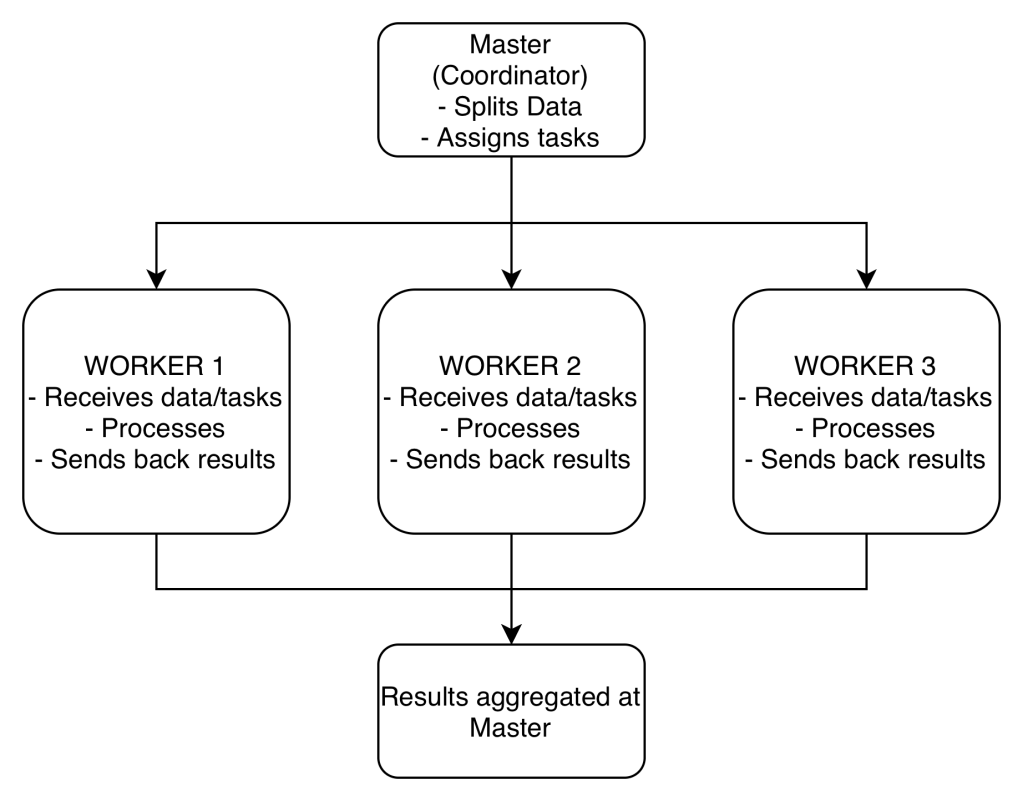
The cluster consists of multiple remote machines connected over a network. One machine acts as the master, and the others act as workers.
- The master manages splitting up the data and the computations.
- The master sends the workers data and calculations to run, and they send their results back to the master.
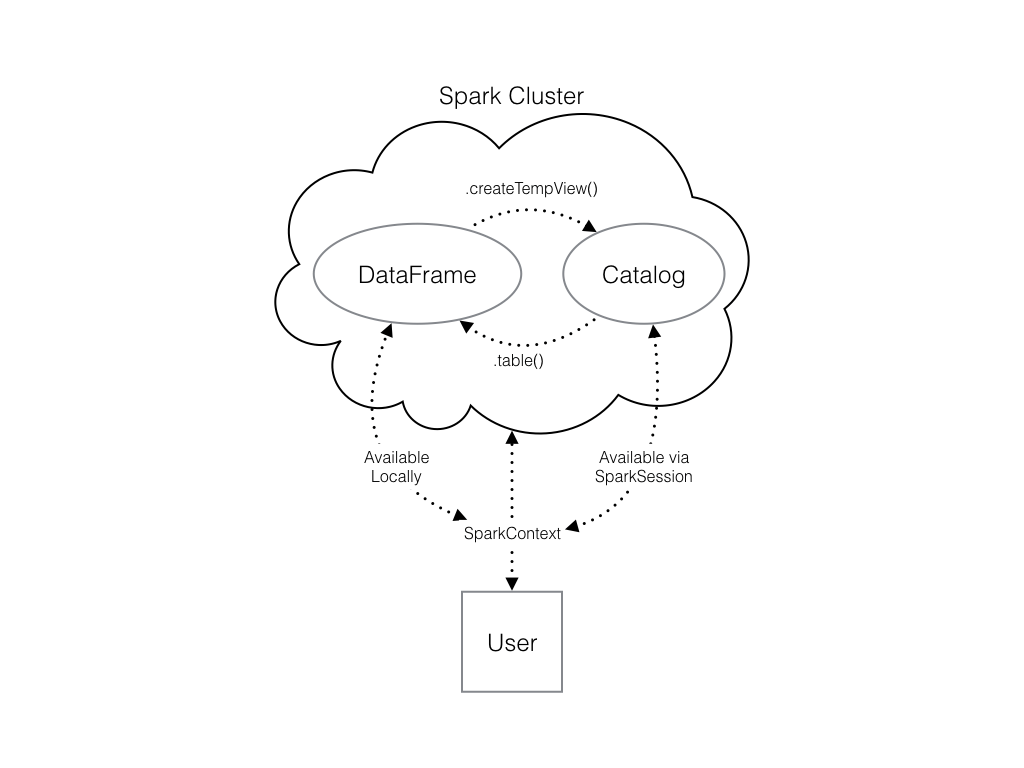
To create a connection, you only need to create an instance of the SparkContext class.
- The class constructor takes a few optional arguments that allow you to specify the attributes of the cluster you’re connecting to.
DataFrames
Spark’s core data structure is the Resilient Distributed Dataset (RDD).
- Low level object that let’s Spark work splitting data across multiple nodes in the cluster.
- RDDs are hard to work with directly, so instead you can use Spark DataFrame abstraction built on top of RDDs.
- The Spark DataFrame was designed to behave a lot like a SQL table (a table with variables in the columns and observations in the rows).
- To start working with Spark DataFrames, you first have to create a
SparkSessionobject from yourSparkContext.- You can think of the
SparkContextas your connection to the cluster and theSparkSessionas your interface with that connection
- You can think of the
In the example below, you get to create a SparkSession, run a query, save the results to a pandas data frame, and print the head of the data frame.
# Import SparkSession from pyspark.sqlfrom pyspark.sql import SparkSession# Create my_sparkmy_spark = SparkSession.builder.getOrCreate()# Print my_sparkprint(my_spark)# Don't change this queryquery = "FROM flights SELECT * LIMIT 10"# Get the first 10 rows of flightsflights10 = my_spark.sql(query)# Show the resultsflights10.show()# Convert the results to a pandas DataFramepd_counts = flight_counts.toPandas()# Print the head of pd_countsprint(pd_counts.head())
It is also possible to put a pandas DataFrame into a Spark cluster. The SparkSession class has a method for this as well.
The .createDataFrame() method takes a pandas DataFrame and returns a Spark DataFrame. The output of this method is stored locally, not in the SparkSession catalog. This means that you can use all the Spark DataFrame methods on it, but you can’t access the data in other contexts.
You can’t use the .sql() method to reference the DataFrame since this will throw an error. You can save the data in a temp table, and use .createTempView() Spark DataFrame method to access the data. You can also use .createOrReplaceTempView() which creates a new temporary table if nothing was there before, or updates an existing table if one was already defined.
# Create pd_temppd_temp = pd.DataFrame(np.random.random(10))# Create spark_temp from pd_tempspark_temp = spark.createDataFrame(pd_temp)# Examine the tables in the catalogprint(spark.catalog.listTables())# Add spark_temp to the catalogspark_temp.createOrReplaceTempView("temp")# Examine the tables in the catalog againprint(spark.catalog.listTables())
You can also load data from external files:
# Don't change this file pathfile_path = "/usr/local/share/datasets/airports.csv"# Read in the airports dataairports = spark.read.csv(file_path, header=True)# Show the dataairports.show()
Add a new column
# Create the DataFrame flightsflights = spark.table("flights")# Show the headflights.show()# Add duration_hrsflights = flights.withColumn("duration_hrs", flights.air_time/60)
Manipulating Data
Filter
# Filter flights by passing a stringlong_flights1 = flights.filter("distance > 1000")# Filter flights by passing a column of boolean valueslong_flights2 = flights.filter(flights.distance > 1000)# Print the data to check they're equallong_flights1.show()long_flights2.show()
Select
# Select the first set of columnsselected1 = flights.select("tailnum", "origin", "dest")# Select the second set of columnstemp = flights.select(flights.origin, flights.dest, flights.carrier)# Define first filterfilterA = flights.origin == "SEA"# Define second filterfilterB = flights.dest == "PDX"# Filter the data, first by filterA then by filterBselected2 = temp.filter(filterA).filter(filterB)# Define avg_speedavg_speed = (flights.distance/(flights.air_time/60)).alias("avg_speed")# Select the correct columnsspeed1 = flights.select("origin", "dest", "tailnum", avg_speed)# Create the same table using a SQL expressionspeed2 = flights.selectExpr("origin", "dest", "tailnum", "distance/(air_time/60) as avg_speed")
Aggregating
# Find the shortest flight from PDX in terms of distanceflights.filter(flights.origin == "PDX").groupBy().min("distance").show()# Find the longest flight from SEA in terms of air timeflights.filter(flights.origin == "SEA").groupBy().max("air_time").show()# Average duration of Delta flightsflights.filter(flights.carrier == "DL").filter(flights.origin == "SEA").groupBy().avg("air_time").show()# Total hours in the airflights.withColumn("duration_hrs", flights.air_time/60).groupBy().sum("duration_hrs").show()
Grouping
# Group by tailnumby_plane = flights.groupBy("tailnum")# Number of flights each plane madeby_plane.count().show()# Group by originby_origin = flights.groupBy("origin")# Average duration of flights from PDX and SEAby_origin.avg("air_time").show()
# Import pyspark.sql.functions as Fimport pyspark.sql.functions as F# Group by month and destby_month_dest = flights.groupBy("month", "dest")# Average departure delay by month and destinationby_month_dest.avg("dep_delay").show()# Standard deviation of departure delayby_month_dest.agg(F.stddev("dep_delay")).show()
Joining
# Examine the dataairports.show()# Rename the faa columnairports = airports.withColumnRenamed("faa", "dest")# Join the DataFramesflights_with_airports = flights.join(airports, on="dest", how="leftouter")# Examine the new DataFrameflights_with_airports.show()
Machine Learning Pipelines
At the core of the pyspark.ml module are the Transformer and Estimator classes.
Transformer classes have a .transform() method that takes a DataFrame and returns a new DataFrame; usually the original one with a new column appended.
Estimator classes all implement a .fit() method. These methods also take a DataFrame, but instead of returning another DataFrame they return a model object. This can be something like a StringIndexerModel for including categorical data saved as strings in your models, or a RandomForestModel that uses the random forest algorithm for classification or regression.
# Rename year columnplanes = planes.withColumnRenamed("year", "plane_year")# Join the DataFramesmodel_data = flights.join(planes, on="tailnum", how="leftouter")# Cast the columns to integersmodel_data = model_data.withColumn("arr_delay", model_data.arr_delay.cast("integer"))model_data = model_data.withColumn("air_time", model_data.air_time.cast("integer"))model_data = model_data.withColumn("month", model_data.month.cast("integer"))model_data = model_data.withColumn("plane_year", model_data.plane_year.cast("integer"))# Create the column plane_agemodel_data = model_data.withColumn("plane_age", model_data.year - model_data.plane_year)# Create is_latemodel_data = model_data.withColumn("is_late", model_data.arr_delay > 0)# Convert to an integermodel_data = model_data.withColumn("label", model_data.is_late.cast("integer"))# Remove missing valuesmodel_data = model_data.filter("arr_delay is not NULL and dep_delay is not NULL and air_time is not NULL and plane_year is not NULL")# Create a StringIndexercarr_indexer = StringIndexer(inputCol="carrier", outputCol="carrier_index")# Create a OneHotEncodercarr_encoder = OneHotEncoder(inputCol="carrier_index", outputCol="carrier_fact")# Create a StringIndexerdest_indexer = StringIndexer(inputCol="dest", outputCol="dest_index")# Create a OneHotEncoderdest_encoder = OneHotEncoder(inputCol="dest_index", outputCol="dest_fact")# Make a VectorAssemblervec_assembler = VectorAssembler(inputCols=["month", "air_time", "carrier_fact", "dest_fact", "plane_age"], outputCol="features")# Import Pipelinefrom pyspark.ml import Pipeline# Make the pipelineflights_pipe = Pipeline(stages=[dest_indexer, dest_encoder, carr_indexer, carr_encoder, vec_assembler])# Fit and transform the datapiped_data = flights_pipe.fit(model_data).transform(model_data)# Split the data into training and test setstraining, test = piped_data.randomSplit([.6, .4])
Model Tuning and Selection
Logistic regression is very similar to a linear regression, but instead of predicting a numeric variable, it predicts the probability (between 0 and 1) of an event
To use this as a classification algorithm, all you have to do is assign a cutoff point to these probabilities. If the predicted probability is above the cutoff point, you classify that observation as a ‘yes’, if it’s below, you classify it as a ‘no’.
A hyperparameter is just a value in the model that’s not estimated from the data, but rather is supplied by the user to maximize performance.
You can tune your logistic regression model using a procedure called k-fold cross validation.
k-fold cross validation
It works by splitting the training data into a few different partitions. The exact number is up to you. Once the data is split up, one of the partitions is set aside, and the model is fit to the others. Then the error is measured against the held out partition. This is repeated for each of the partitions, so that every block of data is held out and used as a test set exactly once. Then the error on each of the partitions is averaged. This is called the cross validation error of the model, and is a good estimate of the actual error on the held out data.
The cross validation error is an estimate of the model’s error on the test set.
# Import the evaluation submoduleimport pyspark.ml.evaluation as evals# Create a BinaryClassificationEvaluatorevaluator = evals.BinaryClassificationEvaluator(metricName="areaUnderROC")# Import the tuning submoduleimport pyspark.ml.tuning as tune# Create the parameter gridgrid = tune.ParamGridBuilder()# Add the hyperparametergrid = grid.addGrid(lr.regParam, np.arange(0, .1, .01))grid = grid.addGrid(lr.elasticNetParam, [0, 1])# Build the gridgrid = grid.build()# Create the CrossValidatorcv = tune.CrossValidator(estimator=lr, estimatorParamMaps=grid, evaluator=evaluator )# Call lr.fit()best_lr = lr.fit(training)# Print best_lrprint(best_lr)
AUC, or area under the curve is a common metric for binary classification algorithms. The curve is the ROC, or receiver operating curve.
The closer the AUC is to one (1), the better the model is.
# Use the model to predict the test settest_results = best_lr.transform(test)# Evaluate the predictionsprint(evaluator.evaluate(test_results))